Just DNA-Seq

Your DNA Guide to a Longer Life
Our reports use detailed genetic data to give you a clear picture of your longevity chances.
Understand What Your Genes Say About Your Health
Find out how your genetic makeup could impact your risk for heart disease and cancer, with clear explanations from trusted sources.


Personalize Your Medicine Cabinet
Discover which drugs work best for you and which to avoid, all based on your unique genetic profile.
Lead a Longer, Healthier Life with Just-DNA-Seq!
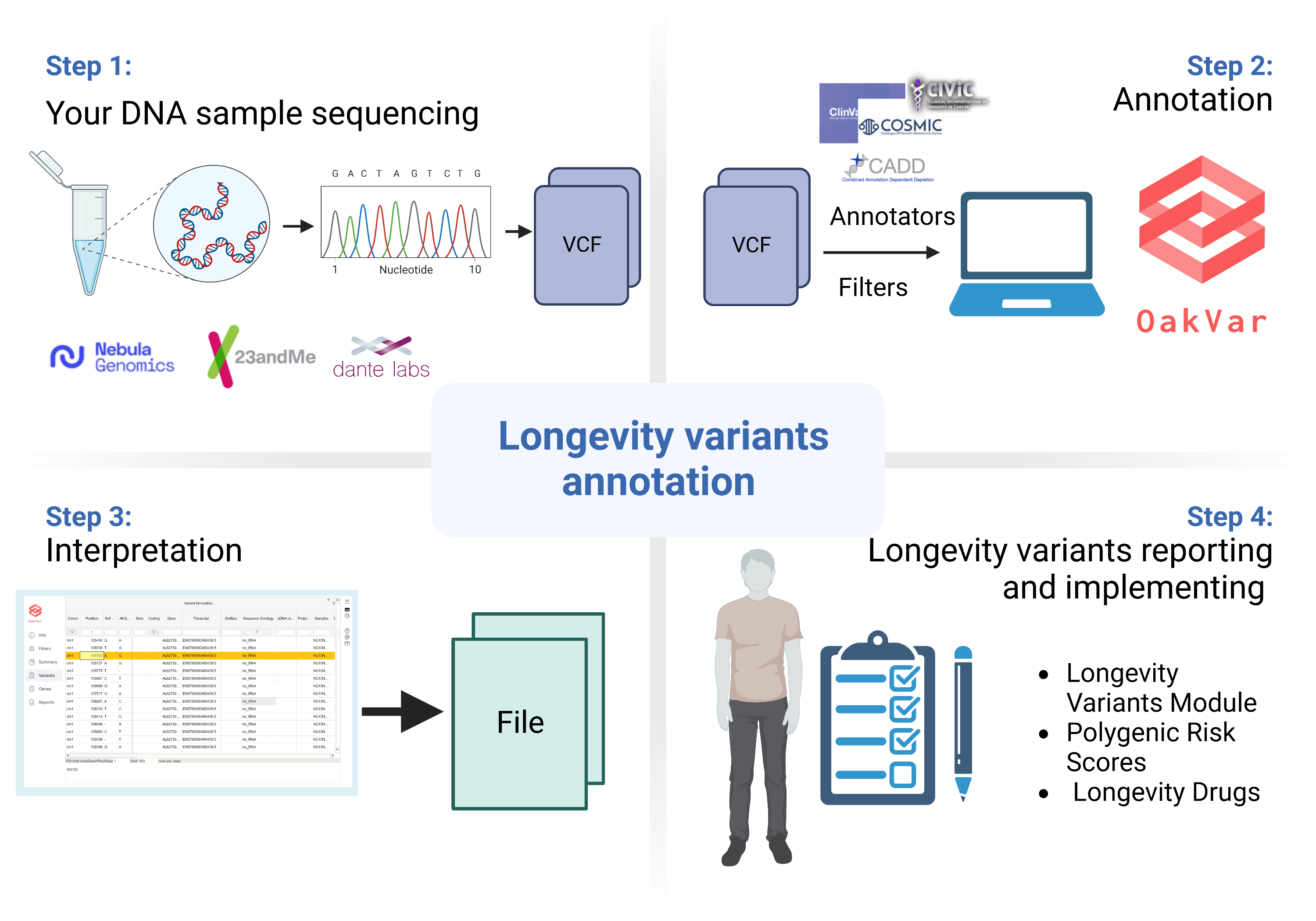
Just-DNA-Seq is an open-source project that helps you analyze your genome:
- Sequence your genome with the help of Nebula, Dante, or any other sequencing company
- Obtain a VCF (Variant Call Format) file containing your genomic data
- Upload this file to our platform for annotation
- Get a personalized report detailing your predisposition to age-related diseases, significant health risks, and insights into your longevity based on your genomic information.
Tutorial on how to use the Just-DNA-Seq platform:
Try Just-DNA-Seq now!
Use our demo account to see already generated reports (login: jdnaseq@gmail.com password: 123456)
At the moment we have the following modules in our report:

|
|
|---|

|
(Currently depricated and removed) Longevity PRS report is based on existing longevity polygenic scores. At the moment, our best performing PRS is implements the score presented in [PMC8087277] and comprises 330 variants. This PRS was shown to be significantly associated with cognitively healthy aging and with prolonged survival. While it is not enough to have “centenarian” genes to become one, they are still needed to cut down major health risks and therefore gain longevity escape velocity. |
|---|

|
|
|---|

|
|
|---|

|
|
|---|

|
|
|---|

|
|
|---|
Our team
- @antonkulaga - bioinformatician at Systems Biology of Aging Group and CellFabrik
- @winternewt - software developer with a chemical background, bioinformatic pipelines developer
- @OlgaBorysova - biologist, geneticist, mitochondria expert, and founder of MitoSpace
- @Alex-Karmazin - senior computer vision engineer, web developer
- @lnalinaf - bioinformatician
- @Lady3mlnm - junior programmer and volunteer at International Longevity Alliance and HEALES
- @marfantyk - software developer with a medical background


